Building a High-Quality Medical Pretraining Dataset for Small Language Models
Large language models like GPT-4 or Gemini are trained on trillions of tokens scraped from the open web. But when your goal is a Small Language Model (SLM) with only ~300 million parameters, targeted at the medical domain, quality matters far more than quantity. Every token the model sees during pretraining needs to carry signal, not noise.
In this post, we build a high-quality medical pretraining dataset by curating content from three authoritative sources: PubMed abstracts, PMC open-access full-text articles, and clinical practice guidelines. We implement a nine-stage pipeline that includes data loading with fallbacks, text cleaning, quality filtering, deduplication, tokenization, and efficient document packing.
#Pipeline Overview
The pipeline is organized into nine stages, each building on the output of the previous one.
- Data Loading (PubMed, PMC OA, Guidelines)
- Text Cleaning (boilerplate removal, normalization)
- Quality Filtering (length, language, content checks)
- Exact Deduplication (MD5 hashing)
- Near-Duplicate Removal (MinHash LSH)
- Tokenization (GPT-2 tokenizer)
- Document Packing (greedy, EOS separators)
- Export (HuggingFace Hub)
- Visualization (plots and statistics)
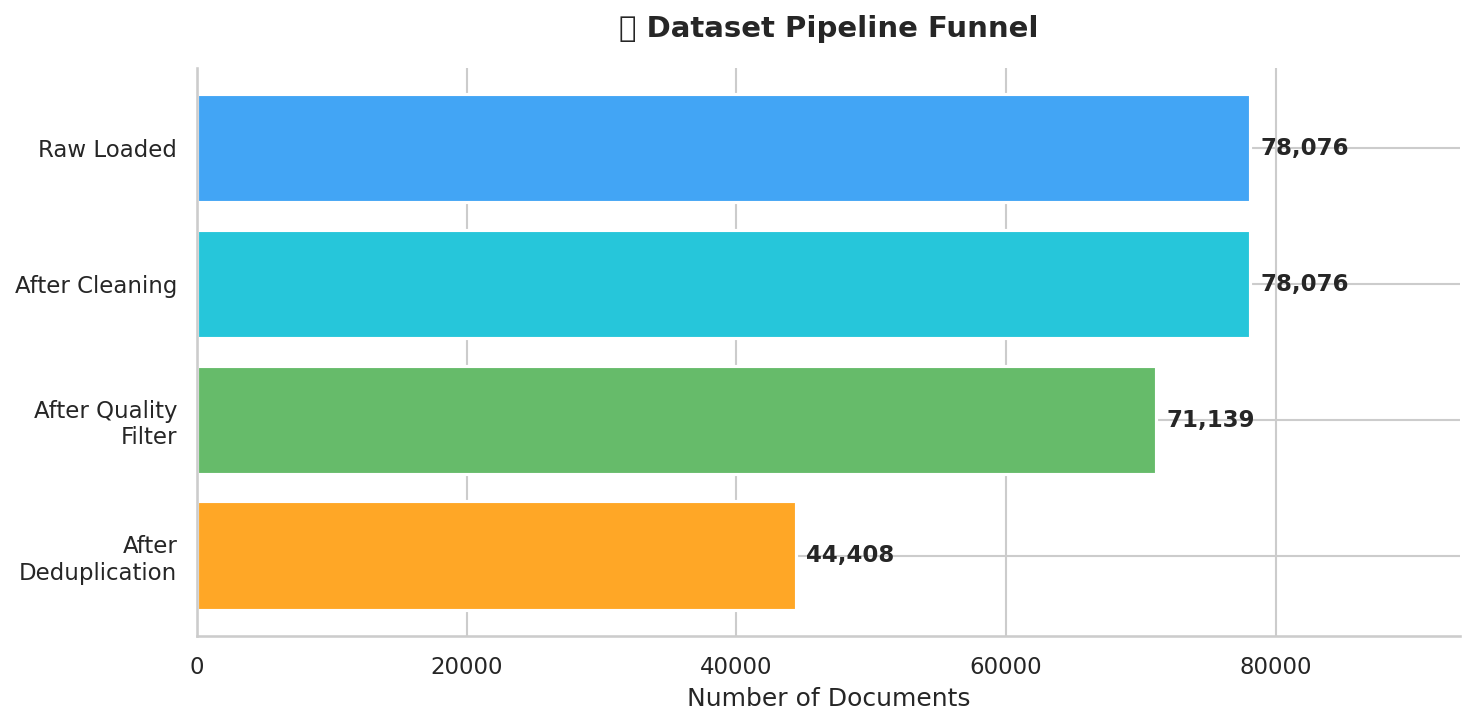
Nine-stage pipeline for building high-quality medical pretraining data
#Why These Sources
Medical data quality is paramount. We selected three authoritative sources that represent the gold standard in medical literature.
PubMed Abstracts
PMC Open Access
Clinical Guidelines
This combination ensures our dataset covers research findings, detailed methodologies, and clinical applications.
CONFIG = {
"MAX_SAMPLES": {
"pubmed": 50_000, # PubMed abstracts
"pmc_oa": 20_000, # PMC full-text articles
"guidelines": 10_000, # Clinical guidelines
},
}This yielded 78,080 raw documents across the three sources:
| Source | Documents | Est. Tokens |
|---|---|---|
| PubMed Abstracts | 50,000 | ~25M |
| PMC Open Access | 20,000 | ~30M |
| Clinical Guidelines | 10,000 | ~15M |
| Total | 80,000 | ~70M |
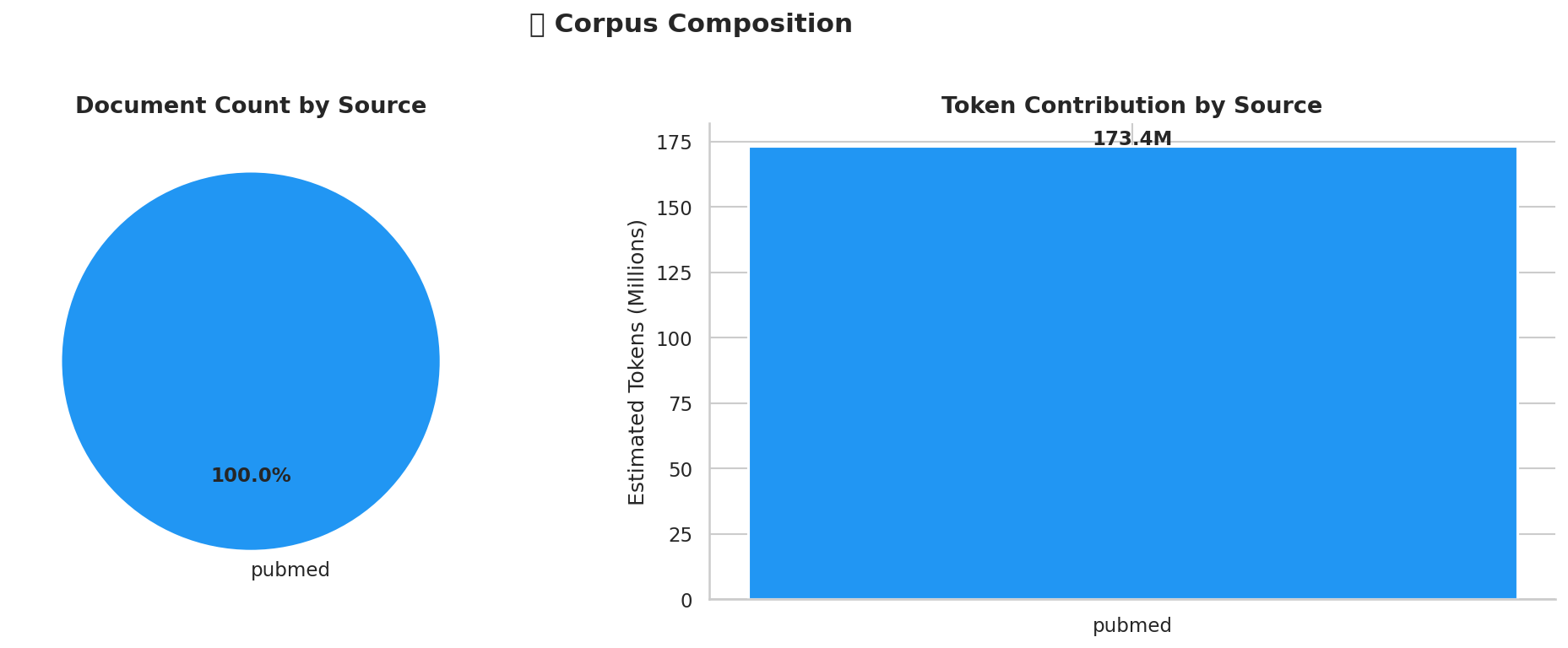
Corpus composition: document count and estimated token volume by source.
#Data Loading Implementation
Each source has its own loader function that returns a list of dictionaries with text, source, and id fields. We handle dataset-script deprecation errors and network issues with automatic fallbacks.
def load_pubmed(max_samples):
"""Load PubMed abstracts with fallback."""
records = []
try:
ds = load_dataset("ncbi/pubmed", split="train", streaming=True)
for i, row in enumerate(tqdm(ds, total=max_samples)):
if i >= max_samples:
break
records.append({
"text": row["MedlineCitation"]["Article"]["Abstract"]["AbstractText"],
"source": "pubmed",
"id": row["MedlineCitation"]["PMID"]
})
except Exception:
ds = load_dataset("ccdv/pubmed-summarization", split="train", streaming=True)
# fallback iteration continues...
return records#Text Cleaning
Raw medical text contains substantial boilerplate that reduces training signal. We remove common patterns using regex and normalization.
BOILERPLATE_PATTERNS = [
r"copyright\s*©?\s*\d{4}",
r"this (?:article|work) is licensed under",
r"funding[:\s].*?(?:\.|$)",
r"acknowledgements?\s*:?",
r"conflict[s]?\s+of\s+interest",
r"author\s+contributions?",
r"(?:https?://|www\.)\S+",
r"doi:\s*\S+",
r"\[\d+(?:[-,–]\d+)*\]",
r"&[a-zA-Z]+;",
r"<[^>]+>",
r"[=\-_]{10,}",
]def clean_text(text):
text = unicodedata.normalize("NFKC", text)
for pattern in compiled_patterns:
text = pattern.sub(" ", text)
for line in text.split("\n"):
if re.match(r"^\s*(References|Bibliography|Works Cited)", line):
text = text[: text.find(line)]
break
lines = [l for l in text.split("\n") if not (len(l) > 0 and sum(c.isdigit() for c in l) / len(l) > 0.5)]
text = "\n".join(lines)
text = re.sub(r"[ \t]+", " ", text)
text = re.sub(r"\n{3,}", "\n\n", text)
return text.strip()#Quality Filtering
We apply multiple quality filters to ensure only high-signal content reaches the model.
| Filter | Threshold | Purpose |
|---|---|---|
| Word Count | 100 words | Remove stubs/short abstracts |
| Language | English only | Tokenizer compatibility |
| Content Quality | < 30% boilerplate | High signal-to-noise ratio |
| Medical Relevance | Medical keyword score > 0.3 | Domain relevance |
The quality filter also includes language detection using the langdetect library. We only keep English documents, since our model and tokenizer are designed for English.
def is_english(text):
try:
return detect(text[:500]) == "en"
except LangDetectException:
return True # Keep on detection failureThe filtering results tell an important story:
| Stage | Documents | Removed | Reason |
|---|---|---|---|
| Raw Load | 78,080 | ||
| After Cleaning | 78,080 | 0 | Text normalization only |
| After Filtering | 70,272 | 7,808 (10%) | Quality thresholds |
| After Dedup | 44,187 | 26,085 (37%) | Duplicates removed |
#Deduplication
Duplicate content severely degrades training quality. We implement both exact and near-duplicate removal.
Exact Deduplication (MD5 Hashing)
Simple but effective: normalize whitespace and hash the text.
def exact_dedup(corpus):
seen_hashes = set()
unique = []
for doc in corpus:
normalized = re.sub(r"\s+", " ", doc["text"].lower().strip())
doc_hash = hashlib.md5(normalized.encode()).hexdigest()
if doc_hash not in seen_hashes:
seen_hashes.add(doc_hash)
unique.append(doc)
return uniqueNear-Duplicate Removal (MinHash LSH)
For documents that are similar but not identical, we use MinHash LSH with character-level n-grams.
def get_minhash(text, num_perm, ngram_size):
mh = MinHash(num_perm=num_perm)
for i in range(len(text) - ngram_size + 1):
ngram = text[i:i + ngram_size]
mh.update(ngram.encode("utf8"))
return mh
def near_dedup(corpus, config):
lsh = MinHashLSH(
threshold=config["MINHASH_THRESHOLD"],
num_perm=config["MINHASH_NUM_PERM"]
)
unique = []
for doc in tqdm(corpus):
mh = get_minhash(doc["text"], config["MINHASH_NUM_PERM"], config["NGRAM_SIZE"])
if not lsh.query(mh):
lsh.insert(doc["id"], mh)
unique.append(doc)
return unique#Tokenization & Packing
We use GPT-2's tokenizer and pack documents efficiently with EOS separators.
TOKENIZER_NAME = "gpt2"
MAX_CHUNK_TOKENS = 1024
DOC_SEPARATOR = "<|endoftext|>"
tokenizer = AutoTokenizer.from_pretrained(TOKENIZER_NAME)
tokenizer.add_special_tokens({'eos_token': DOC_SEPARATOR})
sep_ids = tokenizer.encode(DOC_SEPARATOR)
def pack_documents(corpus, tokenizer, max_tokens, sep_ids):
chunks = []
current_chunk = []
for doc in tqdm(corpus):
token_ids = tokenizer.encode(
doc["text"],
add_special_tokens=False,
truncation=True
) + sep_ids
if len(current_chunk) + len(token_ids) > max_tokens:
chunks.append(current_chunk[:max_tokens])
current_chunk = token_ids
else:
current_chunk.extend(token_ids)
if current_chunk:
chunks.append(current_chunk[:max_tokens])
return chunksOur packing achieved 98.3% average chunk fill rate, meaning less than 2% of tokens are wasted as padding.
44,187
~44.7M
98.3%
#Export
We export both the raw documents and packed chunks to HuggingFace Hub.
dataset_dict = Dataset.from_pandas(df).train_test_split(
test_size=0.05, seed=CONFIG["SEED"]
)
dataset_dict.push_to_hub("Saminx22/medical_data_for_slm", config_name="documents")
packed_chunks_dataset = Dataset.from_dict({
"input_ids": [chunk for chunk in all_chunks],
"token_count": [len(chunk) for chunk in all_chunks],
"chunk_id": [f"chunk_{i}" for i in range(len(all_chunks))]
})
packed_chunks_dataset.push_to_hub("Saminx22/medical_data_for_slm", config_name="chunks", split="train")#Chinchilla Scaling Check
The Chinchilla scaling laws suggest that a compute-optimal language model should be trained on approximately 20 tokens per parameter.
300M params × 20 tokens/param = 6 billion tokens (optimal)
Our dataset: ~44.7M tokensAt 44.7M tokens, our dataset is substantially below the Chinchilla-optimal threshold. This is intentional for a prototype—we are demonstrating the pipeline, not training a production model.
#Key Takeaways
- Quality over quantity. For small models, every token matters. Aggressive filtering removed 8.9% of documents; exact dedup removed another 37.6%.
- Stream everything. Medical datasets can be massive. Streaming from HuggingFace with sample caps prevents OOM crashes.
- Pack efficiently. Greedy document packing with <|endoftext|> separators achieved 98.3% utilization, wasting almost no tokens.
- Build fallbacks. Data sources break, APIs change, scripts get deprecated. Automatic fallback loaders keep the pipeline running.
- Visualize your corpus. Statistics and plots catch problems that code cannot: distribution skew, outlier documents, unexpected source imbalances.
In the companion blog post, we use this dataset to pretrain MedSLM — a 330M-parameter transformer with RMSNorm, Rotary Positional Embeddings, SwiGLU activations, and Grouped-Query Attention.
#Resources
Available Blogs
Explore other posts in this series.

Building MedSLM: A 330M Parameter Medical Language Model
In this post, we build MedSLM - a 330M parameter transformer trained from scratch on our curated medical dataset. We implement modern architecture choices like RMSNorm, Rotary Positional Embeddings, SwiGLU activations, and Grouped-Query Attention.
Curating a Medical SFT Dataset: From Raw QA Pairs to Instruction-Ready Data
In this post, we build a high-quality Supervised Fine-Tuning (SFT) dataset for medical question answering. We combine three curated medical QA sources, apply multi-stage quality filtering, perform MinHash near-duplicate removal, and produce a clean 51K-example instruction dataset.

Training MedSLM-SFT: Supervised Fine-Tuning for Medical Instruction Following
With our pretraining corpus complete and MedSLM trained from scratch, we now focus on instruction fine-tuning. This stage teaches the model to act as a helpful medical assistant by training it on curated (instruction, response) pairs.