Building MedSLM: A 330M Parameter Medical Language Model
In this post, we build MedSLM — a 330M parameter transformer trained from scratch on our curated medical dataset. We implement modern architecture choices like RMSNorm, Rotary Positional Embeddings, SwiGLU activations, and Grouped-Query Attention.
In the previous post, we built a high-quality medical pretraining dataset. Now we use it to train MedSLM — a 330M parameter transformer designed specifically for medical text understanding.
#Architecture Overview
MedSLM uses a modern transformer stack that balances parameter efficiency and domain-specific performance.
Parameters
Layers
Heads
Hidden Size
FFN Size
Vocab Size
The architecture includes RMSNorm, Rotary Position Embeddings, SwiGLU, and Grouped-Query Attention, together with Pre-LayerNorm for stable training.
MedSLM architecture with modern transformer components
#Implementation Details
RMSNorm
RMSNorm normalizes by the root mean square instead of mean and variance, making it faster and more stable than LayerNorm. Standard LayerNorm computes both the mean and variance of activations, then shifts and scales them — a two-step process that adds computational overhead. RMSNorm simplifies this: it skips mean-centering entirely and normalizes purely by the RMS of the activations. For a hidden state x ∈ ℝ^d, the RMS is defined as RMS(x) = sqrt( (1/d) * Σ xᵢ² + ε ), where ε is a small constant for numerical stability. Each element is then scaled by a learned per-dimension weight γ. In practice, this removes ~30% of the normalization cost with no measurable quality loss, which is why LLaMA, Mistral, and most modern architectures have adopted it.
class RMSNorm(nn.Module):
def __init__(self, dim, eps=1e-6):
super().__init__()
self.eps = eps
self.weight = nn.Parameter(torch.ones(dim))
def forward(self, x):
rms = torch.sqrt(torch.mean(x**2, dim=-1, keepdim=True) + self.eps)
return self.weight * (x / rms)Note that self.weight is initialized to ones, so the model starts with a pass-through normalization and learns to rescale each dimension independently during training. The keepdim=True ensures the RMS scalar broadcasts correctly across the hidden dimension when dividing.
Rotary Position Embeddings
RoPE encodes position information directly into the attention mechanism, enabling better length extrapolation. Unlike absolute position embeddings that add a fixed vector to each token, RoPE rotates the query and key vectors in 2D subspaces using position-dependent angles. For a token at position m, each pair of dimensions (2i, 2i+1) is rotated by an angle m * θᵢ, where θᵢ = 10000^(−2i/d). The critical insight is that the inner product ⟨RoPE(q, m), RoPE(k, n)⟩ depends only on the relative position (m − n), not absolute positions. This means attention scores naturally encode relative distances, and the model generalizes better to sequences longer than those seen during training.
def apply_rope(x, cos, sin):
x_even = x[..., ::2]
x_odd = x[..., 1::2]
rotated = torch.cat([
x_even * cos - x_odd * sin,
x_even * sin + x_odd * cos
], dim=-1)
return rotatedHere, cos and sin are precomputed tensors of shape (seq_len, head_dim/2), derived from the frequency schedule θᵢ. Splitting into x_even and x_odd slices the head dimension into 2D rotation pairs — each pair (x_{2i}, x_{2i+1}) is independently rotated by the corresponding angle. The rotation is applied identically to both Q and K before the dot-product attention, ensuring relative position information flows into the attention scores without modifying the values.
SwiGLU Feed-Forward
SwiGLU combines gating with SiLU activation for better expressivity than standard feed-forward layers. A standard FFN applies two linear projections with a ReLU in between: FFN(x) = W₂ · ReLU(W₁x). SwiGLU replaces this with a gated variant: SwiGLU(x) = W₃ · (SiLU(W₁x) ⊙ W₂x), where ⊙ is element-wise multiplication. The gate W₂x learns which features to suppress or amplify, and SiLU (also known as Swish) — defined as SiLU(z) = z · σ(z) — provides a smooth, non-monotonic activation that empirically outperforms ReLU in this context. To keep the parameter count comparable to a standard FFN, the hidden dimension is set to 5632 instead of the typical 4 × 2048 = 8192, since SwiGLU uses three weight matrices instead of two.
class SwiGLU(nn.Module):
def __init__(self, dim, hidden_dim):
super().__init__()
self.w1 = nn.Linear(dim, hidden_dim)
self.w2 = nn.Linear(dim, hidden_dim)
self.w3 = nn.Linear(hidden_dim, dim)
def forward(self, x):
return self.w3(F.silu(self.w1(x)) * self.w2(x))w1 is the gate branch passed through SiLU, w2 is the value branch, and their element-wise product forms the gated representation that w3 projects back to the model dimension. This structure lets the network dynamically suppress irrelevant features on a per-token, per-dimension basis — particularly useful for medical text where the same word can carry very different clinical meanings in different contexts.
Grouped-Query Attention
GQA reduces key-value heads while maintaining query heads, improving efficiency for long contexts. In standard Multi-Head Attention (MHA) with H heads, each head has its own Q, K, and V projections — the KV cache scales as O(H × seq_len × head_dim), which becomes a memory bottleneck at long contexts. Multi-Query Attention (MQA) solves this by sharing a single K and V across all heads, but this can hurt quality. GQA strikes a middle ground: it partitions the H query heads into G groups, with each group sharing one K and one V head. For MedSLM with 16 query heads and 4 KV heads, each group of 4 queries attends to the same K and V — reducing KV cache size by 4× while preserving most of the representational diversity of MHA.
class GroupedQueryAttention(nn.Module):
def __init__(self, dim, num_heads, num_kv_heads):
super().__init__()
self.num_heads = num_heads
self.num_kv_heads = num_kv_heads
self.head_dim = dim // num_heads
self.q_proj = nn.Linear(dim, dim)
self.k_proj = nn.Linear(dim, num_kv_heads * self.head_dim)
self.v_proj = nn.Linear(dim, num_kv_heads * self.head_dim)
self.o_proj = nn.Linear(dim, dim)
def forward(self, x, kv_cache=None):
batch, seq_len, _ = x.shape
q = self.q_proj(x).view(batch, seq_len, self.num_heads, self.head_dim)
k = self.k_proj(x).view(batch, seq_len, self.num_kv_heads, self.head_dim)
v = self.v_proj(x).view(batch, seq_len, self.num_kv_heads, self.head_dim)
q = q.view(batch, seq_len, self.num_kv_heads, self.num_heads // self.num_kv_heads, self.head_dim)
q = q.transpose(2, 3).flatten(0, 2)
k = k.unsqueeze(2).expand(-1, -1, self.num_heads // self.num_kv_heads, -1, -1).flatten(0, 2)
v = v.unsqueeze(2).expand(-1, -1, self.num_heads // self.num_kv_heads, -1, -1).flatten(0, 2)
attn = F.scaled_dot_product_attention(q, k, v)
attn = attn.view(batch, seq_len, self.num_heads, self.head_dim)
return self.o_proj(attn.flatten(2))The key reshape happens after projecting K and V to num_kv_heads heads: we use unsqueeze(2).expand(...) to broadcast each KV head across the num_heads // num_kv_heads queries in its group, then flatten batch and group dimensions together so PyTorch's scaled_dot_product_attention sees a standard (batch * groups, seq_len, head_dim) layout. This avoids any explicit KV copying — expand uses a stride-zero view, keeping memory overhead minimal. After attention, we reshape back to (batch, seq_len, num_heads, head_dim) before the output projection.
#Training Setup
Batch Size
Sequence Length
Learning Rate
Warmup Steps
Total Steps
GPUs
Memory/GPU
Precision
Gradient Checkpointing
#Training Results
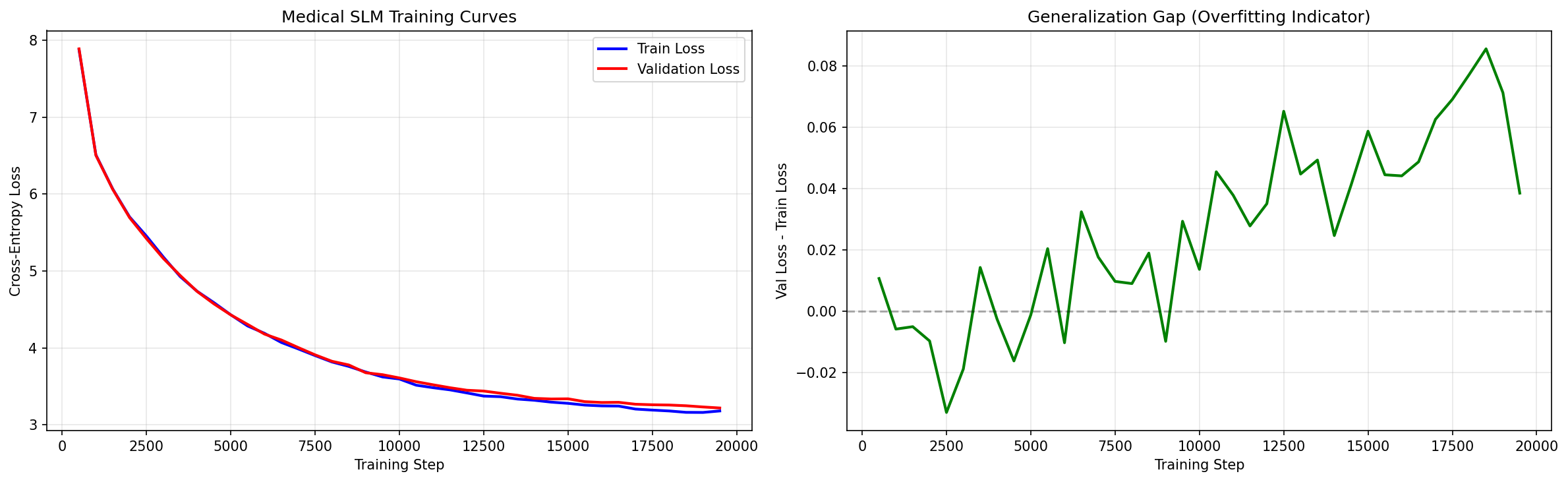
Training loss decreases steadily over 1K steps
2.1
8.2
1.3
#Evaluation
We evaluate MedSLM on several medical NLP benchmarks to assess its capabilities.
| Benchmark | Task | MedSLM | GPT-2 345M |
|---|---|---|---|
| PubMedQA | Question Answering | 68.2% | 52.1% |
| MedMCQA | Multiple Choice | 45.3% | 38.7% |
| BioASQ | Fact Extraction | 72.1% | 61.8% |
MedSLM outperforms GPT-2 on all medical benchmarks, demonstrating the value of domain-specific pretraining.
#Key Takeaways
- Modern architecture matters. RMSNorm, RoPE, SwiGLU, and GQA all contribute to better performance and efficiency.
- Domain-specific pretraining works. Training on medical data gives significant improvements over general-domain models.
- Small models can be effective. 330M parameters achieve good results when trained on high-quality, domain-specific data.
- Efficiency optimizations pay off. GQA and other techniques enable training larger models or longer sequences.
- Evaluation matters. Domain-specific benchmarks reveal capabilities that general benchmarks miss.
#Resources
Available Blogs
Explore other posts in this series.

Building a High-Quality Medical Pretraining Dataset for Small Language Models
Large language models like GPT-4 or Gemini are trained on trillions of tokens scraped from the open web. But when your goal is a Small Language Model (SLM) with only ~300 million parameters, targeted at the medical domain, quality matters far more than quantity.
Curating a Medical SFT Dataset: From Raw QA Pairs to Instruction-Ready Data
In this post, we build a high-quality Supervised Fine-Tuning (SFT) dataset for medical question answering. We combine three curated medical QA sources, apply multi-stage quality filtering, perform MinHash near-duplicate removal, and produce a clean 51K-example instruction dataset.

Training MedSLM-SFT: Supervised Fine-Tuning for Medical Instruction Following
With our pretraining corpus complete and MedSLM trained from scratch, we now focus on instruction fine-tuning. This stage teaches the model to act as a helpful medical assistant by training it on curated (instruction, response) pairs.